Genetic contribution to psychiatric disorders
The substantial influence of genetic variation on risk for a broad range of psychiatric disorders has been initially estimated from twin studies and more recently directly assessed from large-scale genomic studies.1
Psychiatric disorders are highly polygenic, with common variations accounting for a large portion of heritability. Genome-wide association studies (GWASs) of the past decade have allowed a new approach to understanding the genetic contribution to and similarities between psychiatric disorders.2 Many risk loci have emerged from these GWAS, among others, schizophrenia (SCZ), bipolar disorder (BD), major depressive disorder (MDD), and attention-deficit/hyperactivity disorder (ADHD).3 A surprisingly high degree of genetic overlap has been shown among psychiatric disorders.4 Furthermore, this genetic overlap may extend to other medical conditions, including cardiometabolic diseases, indicating that genetic vulnerability is rarely specific. Testing the borders of the genetic vulnerability of psychiatric conditions, such as MDD, is therefore fascinating, as it shows commonalities and differences with other psychiatric and somatic conditions or even risk factors, challenging our understanding of the concept of depression.
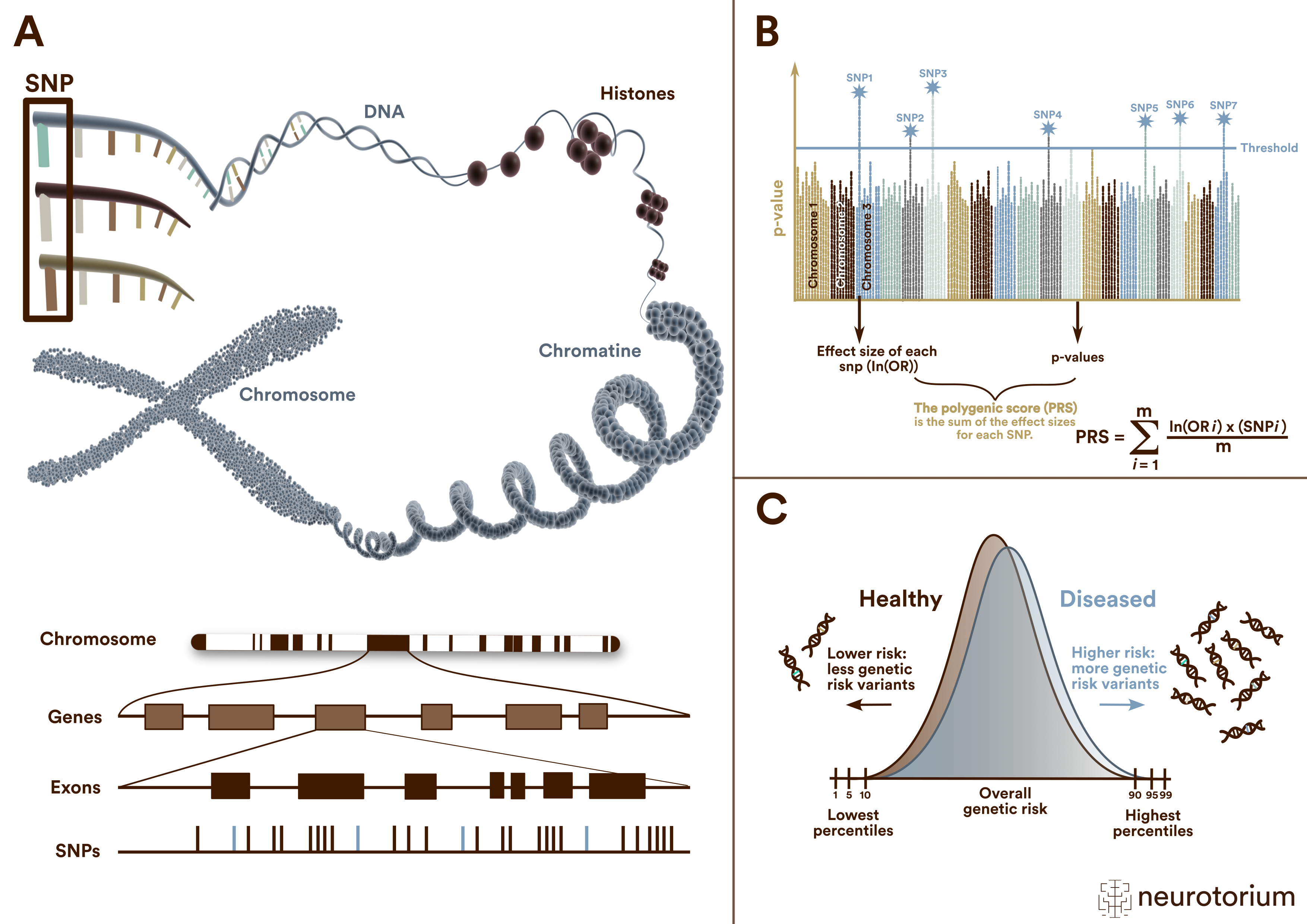
Figure 1: Single nucleotide polymorphisms (SNPs), Polygenic risk scores (PRS) and their distribution
(A) A single nucleotide polymorphism (SNP) is a frequent (>1%) genomic variant at a single base position in the DNA. Some SNPs in the coding region of a gene (exon) change the amino acid sequence of a protein, and others in the coding region do not affect the protein sequence.
(B) Polygenic risk scores (PRS) are typically calculated by summing the weighted allele counts of independent (i.e., uncorrelated) SNPs found to be associated with a trait or disorder in a genome-wide association study (GWAS). For quantitative traits, the weights are the linear regression effect sizes, i.e., the β coefficients. The weights for case/control phenotypes are the natural logarithm of the odds ratio, ln(OR).
(C ) Polygenic risk scores (PRS) follow a normal distribution on a population level. The distributions of cases and controls clearly overlap. Thus, meaningful risk predictions can be only expected for extreme quantiles at the top and bottom of the PRS distributions.
h2SNP, PRS, and their relevance in genetic research in psychiatry
Heritability (h2) is the proportion of the variation in liability to a disease that can be attributed to genetic factors and is calculated based on the inference of genetic factors shared between relatives.2 Estimation of h2 previously relied on closely related individuals, such as siblings or twins. However, recent advancements in genetic research have introduced the concept of single nucleotide polymorphism (SNP)-based heritability (h2SNP), which estimates heritability using single nucleotide polymorphisms (SNPs) across the entire genome of unrelated individuals.
Using h2SNP allows for a more comprehensive understanding of the genetic architecture of psychiatric disorders and provides valuable insights into the shared genetic factors underlying these conditions and others. h2SNP does not involve de novo mutations, though, explaining why h² (the global estimated heritability) is always larger than h²SNP (the part of heritability attributed to SNPs only). Furthermore, h2SNP specifically refers to common DNA variants with an allele frequency over 1%.
The “missing heritability” concept is directly derived from the difference between h² and h2SNP. The ratio of SNP-based to total heritability describes the relative importance of common variants to the genetic architecture, which could differ between disorders.5 Estimates of heritability and SNP-based h2 provide guidelines for maximum future accuracy of risk prediction. Genetic relations between psychiatric disorders and nonpsychiatric disorders can also be derived from SNP-based genetic correlations. These correlations can reflect pleiotropy and also causal relationships.
Polygenic risk scores (PRS) are predictors of genetic susceptibility to diseases, calculated for individuals as weighted counts of millions of variants, in which the risk variants and their weights have been identified in GWAS. Its formula, although adding millions of parameters, is therefore easy to compute, where PRS corresponds to the sum of the OR for each variant divided by the number of these variants.
Recent advancements in genetic research have introduced the concept of single nucleotide polymorphism (SNP)-based heritability, which estimates heritability using single nucleotide polymorphisms (SNPs) across the entire genome of unrelated individuals.
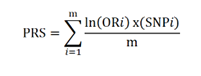
PRS are promising tools for aiding clinical decision-making in many areas of medical practice, including psychiatry.6 Like h2SNP, PRS only capture part of the genetic contribution to psychiatric diseases and cannot be used alone to predict complex diseases like mental health disorders. Combining PRS with other risk factors (e.g., childhood traumas) may improve the prediction of different outcomes, such as adverse physical health issues or response to treatment in clinical populations.
PRS are promising tools for aiding clinical decision-making in many areas of medical practice, including psychiatry.6 Like single nucleotide polymorphism (SNP)-based heritability, PRS only capture part of the genetic contribution to psychiatric diseases and cannot be used alone to predict complex diseases like mental health disorders.
The relevance of PRS and h2SNP in genetic research of MDD
The genetic basis of MDD
Heritability for major depression is around 50%, but the one based on single nucleotide polymorphisms (h2SNP) was estimated to be 8.9% on the liability scale, based on a lifetime risk of 0.15.4 The most recent published GWAS for major depression, including 246,363 cases and 561,190 controls, identified 102 loci significantly associated with MDD, highlighting the highly polygenic nature of the disease.4
The inconsistency in genetic results in MDD is likely due to differences in methodological approaches or in summary statistics but could also be due to the clinical (phenotypic) heterogeneity of MDD. MDD onset can occur at any stage of life, but the factors associated with MDD are often age-specific or age-restricted.7 For instance, increased genetic risk for MDD is associated with earlier age at onset (AAO) compared with later.3 Earlier AAO MDD has a higher heritability and is associated with increased risk for MDD in relatives. On the other hand, vascular disease and its risk factors are linked to a later AAO for MDD.8,9,10 A large study of Swedish twins showed that a later AAO for MDD in one twin was associated with a higher risk for vascular disease in the other twin.11
The most recent published GWAS for major depression, including 246,363 cases and 561,190 controls, identified 102 loci significantly associated with MDD, highlighting the highly polygenic nature of the disease.
A better understanding of the links between MDD and other psychiatric conditions
Figure 2 represents the SNP-based genetic correlations between eight major psychiatric disorders, including MDD. They were depicted using an indirect graph to reveal complex genetic relationships.
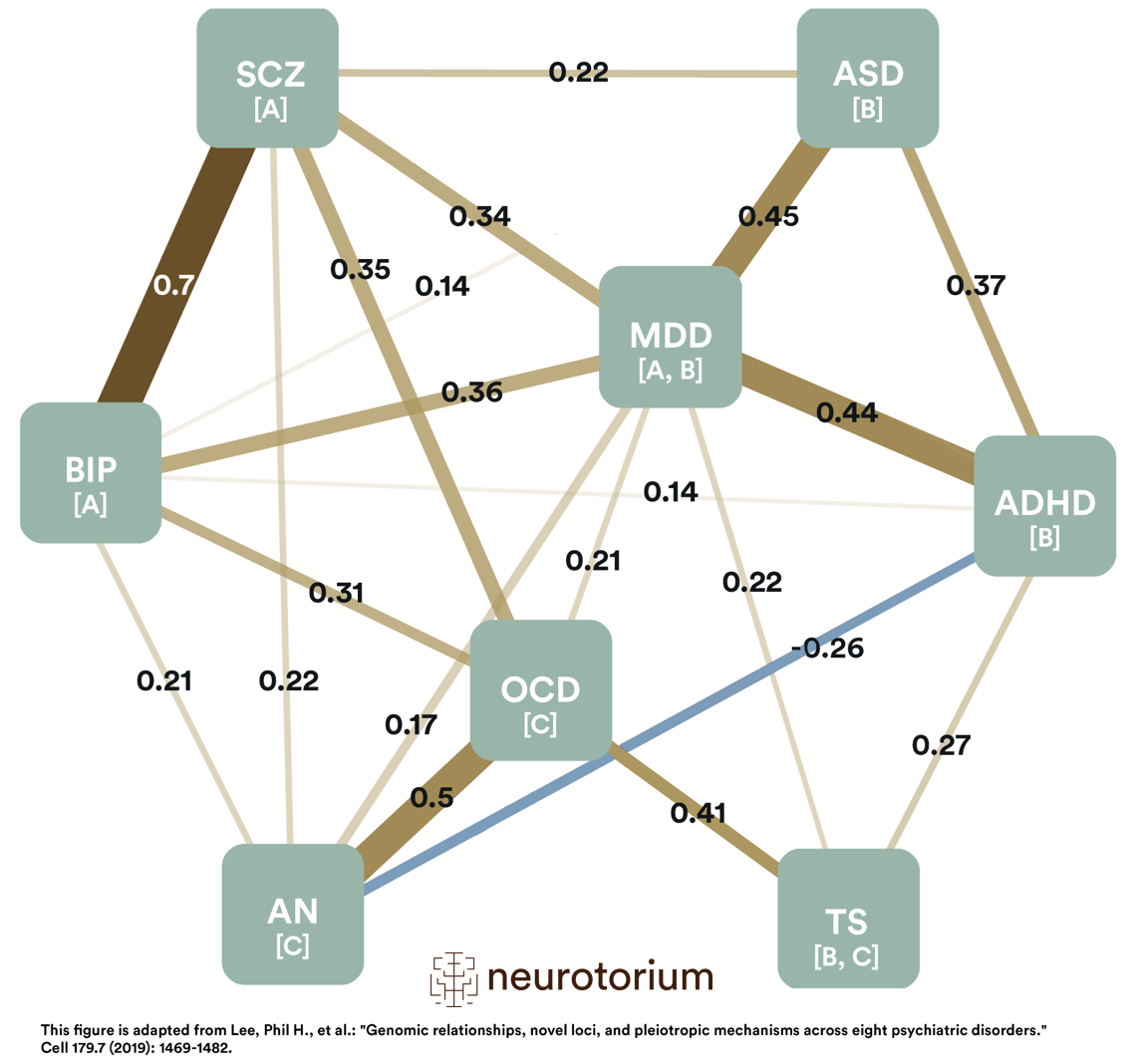
Figure 2: Single nucleotide polymorphisms (SNP)-based genetic correlations between eight major psychiatric disorders
ADHD: Attention-Deficit/Hyperactivity Disorder; AN: Anorexia Nervosa; ASD: Autism Spectrum Disorder; BIP: Bipolar Disorder; MDD: Major Depressive Disorder; OCD: Obsessive-Compulsive Disorder; SCZ: Schizophrenia; TS: Tourette Syndrome.
This figure represents the SNP (Single Nucleotide Polymorphism)-based genetic correlations between 8 major psychiatric disorders. Each node represents a disorder, with numbers on the lines indicating the strength of the pairwise correlations. The width of the lines increases with the absolute values of genetic correlations (rg). The first group of disorders is annotated with “A”; these are the mood and psychotic disorders. The second group, represented with “B,” is early-onset neurodevelopmental disorders. The third group is marked with “C,” disorders with compulsive behaviors. Please note that both MDD and TS belong to two groups. While MDD belongs to mood and psychotic disorders (A) and early-onset neurodevelopmental disorders (B), TS belongs to early-onset neurodevelopmental disorders (B) and disorders with compulsive behaviors (C). This figure is adapted from Lee, Phil H., et al.: “Genomic relationships, novel loci, and pleiotropic mechanisms across eight psychiatric disorders.” Cell 179.7 (2019): 1469-1482.
The modeling of genetic correlations among the eight studied disorders using two different methods identified three groups of disorders based on shared genomics: one consisting of disorders characterized by compulsive behaviors (anorexia nervosa, obsessive-compulsive disorder, and Tourette syndrome), a second consisting of mood and psychotic disorders (MDD, bipolar disorder, and schizophrenia), and a third composed of early-onset neurodevelopmental disorders (Autism spectrum disorder, attention-deficit/hyperactivity disorder, MDD and Tourette syndrome).
These findings show discrepancies between the biological underpinnings of psychiatric diseases and how we currently classify them clinically. These results conflict with the traditional, categorical definition of mental disease.1 The loading of MDD on two factors may indicate biological heterogeneity within MDD, which is in line with recent research demonstrating a link between early-onset depression and phenotypes related to neurodevelopment and genetic risk for ADHD.12
The loading of MDD on two factors may indicate biological heterogeneity within MDD, which is in line with recent research demonstrating a link between early-onset depression and phenotypes related to neurodevelopment and genetic risk for ADHD.
A better understanding of the links between MDD and other medical conditions
Non-psychiatric comorbidities are very common in patients with psychiatric diseases. These comorbidities might rather be co-occurrences underlying a global systemic disorder with shared etiopathogenic mechanisms. A recent study explored whether the association between MDD and cardio-metabolic traits is partly due to genetic factors.13 Using data from GWAS that was available to the public as well as from the PGC, UK Biobank, and Generation Scotland, authors demonstrate a genetic overlap between MDD and cardio-metabolic traits (body mass index (BMI), coronary artery disease, type 2 diabetes) based on genetic correlations of the PRS (respectively, rg = 0.13, p = 3.43 × 10−11), rg = 0.12, p = 8.47 × 10−6) and rg = 0.11, p = 0.0001).
These correlations were largely irrespective of AAO for MDD, particularly for coronary artery disease and type 2 diabetes. The association with BMI showed evidence of a stronger link with later AAO MDD. Cardio-metabolic traits and their risk factors are open to preventative strategies; therefore, further understanding of the shared genetic aetiology between cardio-metabolic traits and MDD could play an important role in preventing or managing MDD.
Other medical conditions also share a genetic component with MDD, such as osteoarthritis,14 atopic diseases (asthma, hay fever, eczema)15 or even migraine.16 Deciphering the genetic relationships between MDD and other medical conditions may facilitate understanding of their biological mechanisms and developing novel treatment approaches.
Cardio-metabolic traits and their risk factors are open to preventative strategies; therefore, further understanding of the shared genetic aetiology between cardio-metabolic traits and MDD could play an important role in preventing or managing MDD.
A better understanding of intermediate factors
Genes code for proteins and enzymes; therefore the depressive episode can be considered as the endpoint of a probably complex cascade. Confronting the phenotype to PRS can be used to detect more closely related intermediate factors. For instance, a robust sex-specific main effect of the PRS of MDD on bilateral infant caudate volumes was observed in children of depressed mothers.17 This sex-specific polygenic effect might lay the ground for the higher prevalence of depression in women compared to men.
In a recent study on blood markers and PRS in MDD, a large proportion of marker associations for MDD PRS were immune- and inflammatory-related, supporting previous works showing that aberrant immune and inflammatory processes contribute to and possibly underlie this disorder, influenced by genetic risk and robust to controlling for lifestyle factors.18 Interestingly, the genetic liability for major depression, indexed par MDD PRS, also increases the risk for suicide attempts across psychiatric disorders (SCZ, MDD, BD).19 Individuals with elevated polygenic risk for MDD may also be at risk for alcohol dependence.20
In a recent study on blood markers and PRS in MDD, a large proportion of marker associations for MDD PRS were immune- and inflammatory-related, supporting previous works showing that aberrant immune and inflammatory processes contribute to and possibly underlie this disorder, influenced by genetic risk and robust to controlling for lifestyle factors.
A clinician help decision predicting score
Differentiation between MDD and other psychiatric conditions such as anxious disorders or bipolar disorder (BD) can be clinically challenging while it has important prognostic and therapeutic implications. In a recent study, authors tested whether a PRS improve clinically based classification models of BD versus MDD diagnosis.21 They found a utility of PRS in subjects at the extremes of the distribution and/or subjects for whom clinical symptoms were poorly measured or yet to manifest.Another study demonstrates that PRS effectively predicts depression onset in Alzheimer’s disease based on MDD risk variants.22
Overall, the use of PRS in MDD detection might be constrained by its uncertainty. One promising way to further extend its usability is fusion with other biomarkers (magnetic resonance imaging, inflammatory markers, or voice). A study constructed an MDD biomarker by combining the PRS and voice features with promising results toward this approach.23
A potential novel approach for therapeutics of depressive disorders
Improving response and remission rates in MDD remains a significant challenge, as well as predicting propensity to side effects. Matching patients to the treatment they will most likely respond to should be the ultimate goal. Even though numerous studies have investigated patient-specific indicators of treatment efficacy, there are no (bio)markers or empirical tests for use in clinical practice. However, there has been evidence of a complex yet clear genetic architecture of antidepressant response with PRS involved in antidepressant responses. A recent systematic review demonstrated the pleiotropy between antidepressant treatment response and other traits. For instance, PRS for ADHD, the personality trait openness, but also coronary artery disease, obesity, and stroke have been associated with antidepressant treatment response in patients with unipolar MDD.24
Using seasonality and clock genes PRS might also be a promising strategy.25 A recent study demonstrated a polygenic overlap between loneliness, severe mental disorders, and cardiovascular disease risk factors, suggesting shared molecular mechanisms and potential common therapeutic targets.26
Finally, treatment-resistant depression probably represents a specific phenotype among MDD, with distinct genetic and clinical-demographic characteristics compared with non-resistant depression. Fabbri and colleagues demonstrated on a large sample a different genetic architecture (underpinned by a different h2SNP) between non-resistant and resistant MDD, with the latter sharing a strong genetic predisposition with ADHD, which corroborates clinical findings.27 PRS will allow a more dimensional approach by elucidating the mechanisms of common phenotypes or dimensions across diseases, in line with Research Domain Criteria (RDoC) approaches.
A recent study demonstrated a polygenic overlap between loneliness, severe mental disorders, and cardiovascular disease risk factors, suggesting shared molecular mechanisms and potential common therapeutic targets.
Conclusion
As clinical psychiatry develops to incorporate data from molecular, clinical, environmental, and lifestyle metrics, polygenic risk scores can contribute to risk assessment and outcome prediction. Over the last few years, the decreased costs and enhanced accessibility to large GWAS datasets have laid the foundations for developing PRS and h2SNP evaluations. Expectations of what PRS can and cannot provide should be considered when evaluating the utility of PRS in clinical psychiatry, as well as ethical concerns related to their use. The future of PRS and h2SNP includes larger datasets, better definition of phenotypes, and so-called comorbidities or co-occurrences (severity, age at onset, and other conditions).
The future of PRS and single nucleotide polymorphism (SNP)-based heritability includes larger datasets, better definition of phenotypes, and so-called comorbidities or co-occurrences (severity, age at onset, and other conditions).






